Table of Contents
Python Image Processing Cookbook
Loading Image File
matplotlib has matplotlib.image.imread, and PIL also has similar file input methods. The former is limited to png files and the latter is limited to 8-bit images. For this reason, we focus on openCV and TiffFile modules.
openCV
Tiff file to numpy.ndarray
>>> import cv
>>> img = cv.imread('/Users/miura/img/blobs.tif')
>>> type(img)
>>> <type 'numpy.ndarray'>
- cv.imread reads only the first frame in tiff stack.
- 8bit image becomes RGB-like format, triplet of 8 bits per pixel.
Tiff file to Iplimage
In [40]: img = cv.LoadImage('/Users/miura/img/blobs.tif')
In [41]: img
Out[41]: <iplimage(nChannels=3 width=256 height=254 widthStep=768 )>
Tiff file to cvMat
Tifffile.py
Tiff to numpy.ndarray, Single Image
In [44]: import tifffile as tff
In [50]: tiffimg = tff.TIFFfile('/Users/miura/img/blobs.tif')
In [51]: type(tiffimg)
Out[51]: <class 'tifffile.TIFFfile'>
In [52]: img = tiffimg.asarray()
In [53]: type(img)
Out[53]: <type 'numpy.ndarray'>
Tiff to numpy.ndarray, Stack Image
In [54]: tiffimg = tff.TIFFfile('/Users/miura/img/flybrainG.tif')
In [55]: img = tiffimg.asarray()
In [57]: tiffimg
Out[57]: <tifffile.TIFFfile object at 0x1299ac910>
In [65]: type(img)
Out[65]: <type 'numpy.ndarray'>
In [61]: img10 = tiffimg[10].asarray()
In [62]: type(img10)
Out[62]: <type 'numpy.ndarray'>
- if the image file is a single frame image, not so different from the others
- Stack tiff file is loaded peroperly. Single frame is extractable by indexing. In above case, 11th frame is extracted.
Conversion between types
OpenCV Mat object to numpy.ndarray object
- tested with python2.6, openCV 2.2, numpy 1.6.1, OSX10.6.8
>>> import cv
>>> import numpy as np
>>> mat = cv.CreateMat( 3 , 5 , cv.CV_32FC1 )
>>> cv.Set( mat , 7 )
>>> a = np.asarray( mat[:,:] )
>>> a
array([[ 7., 7., 7., 7., 7.],
[ 7., 7., 7., 7., 7.],
[ 7., 7., 7., 7., 7.]], dtype=float32)
<link>
OpenCV Image object to numpy.ndarray object
- tested with python2.6, openCV 2.2, numpy 1.6.1, OSX10.6.8
>>> im = cv.CreateImage( ( 5 , 5 ) , 8 , 1 )
>>> cv.Set( im , 100 )
>>> im_array = np.asarray( im )
>>> im_array
array(<iplimage(nChannels=1 width=5 height=5 widthStep=8 )>, dtype=object)
>>> im_array = np.asarray( im[:,:] )
>>> im_array
array([[100, 100, 100, 100, 100],
[100, 100, 100, 100, 100],
[100, 100, 100, 100, 100],
[100, 100, 100, 100, 100],
[100, 100, 100, 100, 100]], dtype=uint8)
<link>
Data Visualization
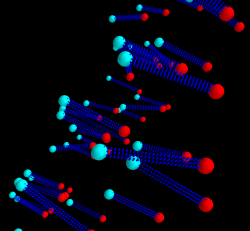
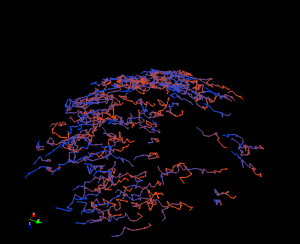
plot 3D coordinates using Mayavi
An example script for loading (x, y, y) coordinates data from tab-delimited text file and plot them in 3D using mayavi2.
from matplotlib import mlab as matp
filename = '/Users/miura/data.txt'
x1, y1, z1, x2, y2, z2 = matp.load(filename, usecols=[0, 1, 2, 3, 4, 5], unpack=True)
from mayavi.mlab import points3d
from mayavi.mlab import plot3d
from mayavi import mlab as maya
p1s = points3d(x1, y1, z1, scale_factor=.25, color=(0, 1, 1))
p2s = points3d(x2, y2, z2, scale_factor=.25, color=(1, 0, 0))
for idx, xval in enumerate(x1):
plin1 = plot3d([x1[idx], x2[idx]], [y1[idx], y2[idx]], [z1[idx], z2[idx]], tube_radius=0.1, colormap='Spectral', color=(0, 0, 1))
maya.show()
In this example, we assume the follwoing data structure in the file: a pair of coordinates per line, so 6 numbers are in one line separated by tab. It should look like
1.0 3.6 4.8 5.1 6.12 7.14 ...
To run this script, the best is to run it from ipython with thread so first you start ipython by
ipython -wthread
or incase of newer ipython (version >= 1.1)
ipython --i
then in the ipython interface,
run exampleMayavi.py
A new window pops up, and after drawing of the scene is finished, you could control the scene by such as
maya.view(100, 40)
or to animate the scene
for i in range (1, 360, 3): maya.view(i, i)
to close the scene,
maya.close()
Note that in this example,
from mayavi import mlab as maya
since the namespace “mlab” overlaps with matplotlib.mlab.